In the cutting-edge research lab at Helix Institute of Molecular Biology, scientists are analyzing strands of DNA to understand how nucleotides pair under spatial constraints.
You are part of a bioinformatics team tasked with analyzing a DNA sequence represented as a string of length , composed of the nucleotides:
- Adenine ()
- Thymine ()
- Cytosine ()
- Guanine ()
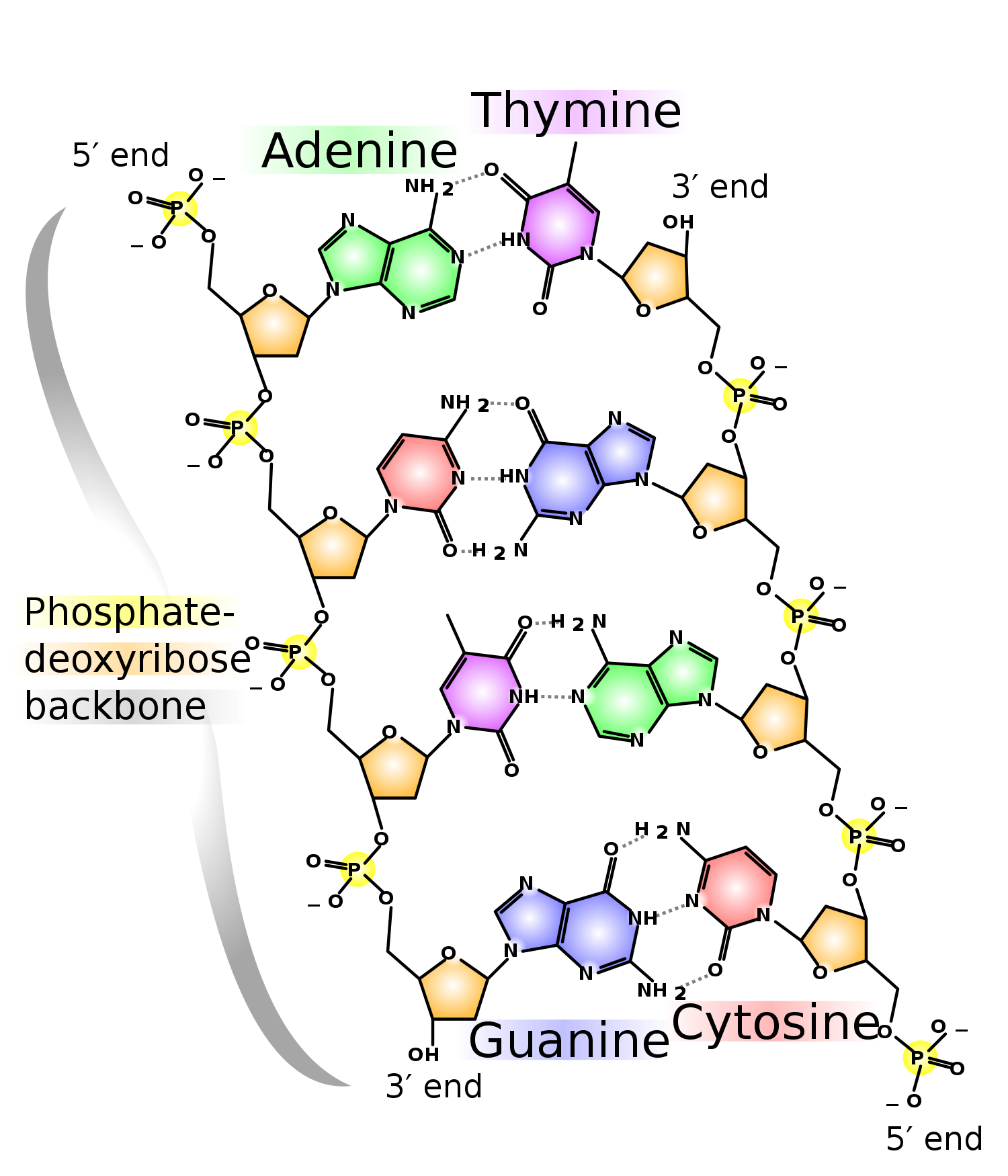
As per Watson-Crick base pairing rules, pairs with , and pairs with .
Task
You are given a string of length , made up of characters , , and . We define the distance between the characters at positions and as being . You are also given queries of the form , where you have to count the number of pairs, such that they are at a distance between and in the string.
Input data
The input file consists of:
- a line containing integers , .
- a line containing the character string .
- lines, the -th of which consisting of the integers .
Output data
The output file must contain the answer for each query on a separate line.
Constraints and clarifications
- .
- .
- .
- for each query.
- and are considered valid pairs.
| # | Score | Constraints |
|---|---|---|
| 1 | 0 | Examples |
| 2 | 10 | |
| 3 | 5 | |
| 4 | 15 | |
| 5 | 10 | |
| 6 | 15 | |
| 7 | 45 | No additional restrictions |
Example
stdin
8 8
ATCGCGAT
1 1
2 2
3 3
4 4
5 5
6 6
7 7
1 7
stdout
5
0
1
0
1
0
1
8
Explanation
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .
- The valid pairs of distance are the following: .